AMBIC Data Platform
Full-stack web platform for biomanufacturing data: ingestion, normalization, visualization, and sharing across research teams. Handles life-critical data with provenance tracking and granular access control.
Context
Researchers at the Advanced Mammalian Biomanufacturing Innovation Center (AMBIC) work with process data in CSV, XLSX, and JSON from a range of instruments and equipment. Before this platform, data sharing between teams meant emailing spreadsheets, and there was no standardized schema across datasets.
Because this data feeds into pharmaceutical manufacturing decisions, integrity requirements are high. A normalization bug that silently drops rows or misaligns columns could lead to incorrect conclusions in a clinical context.
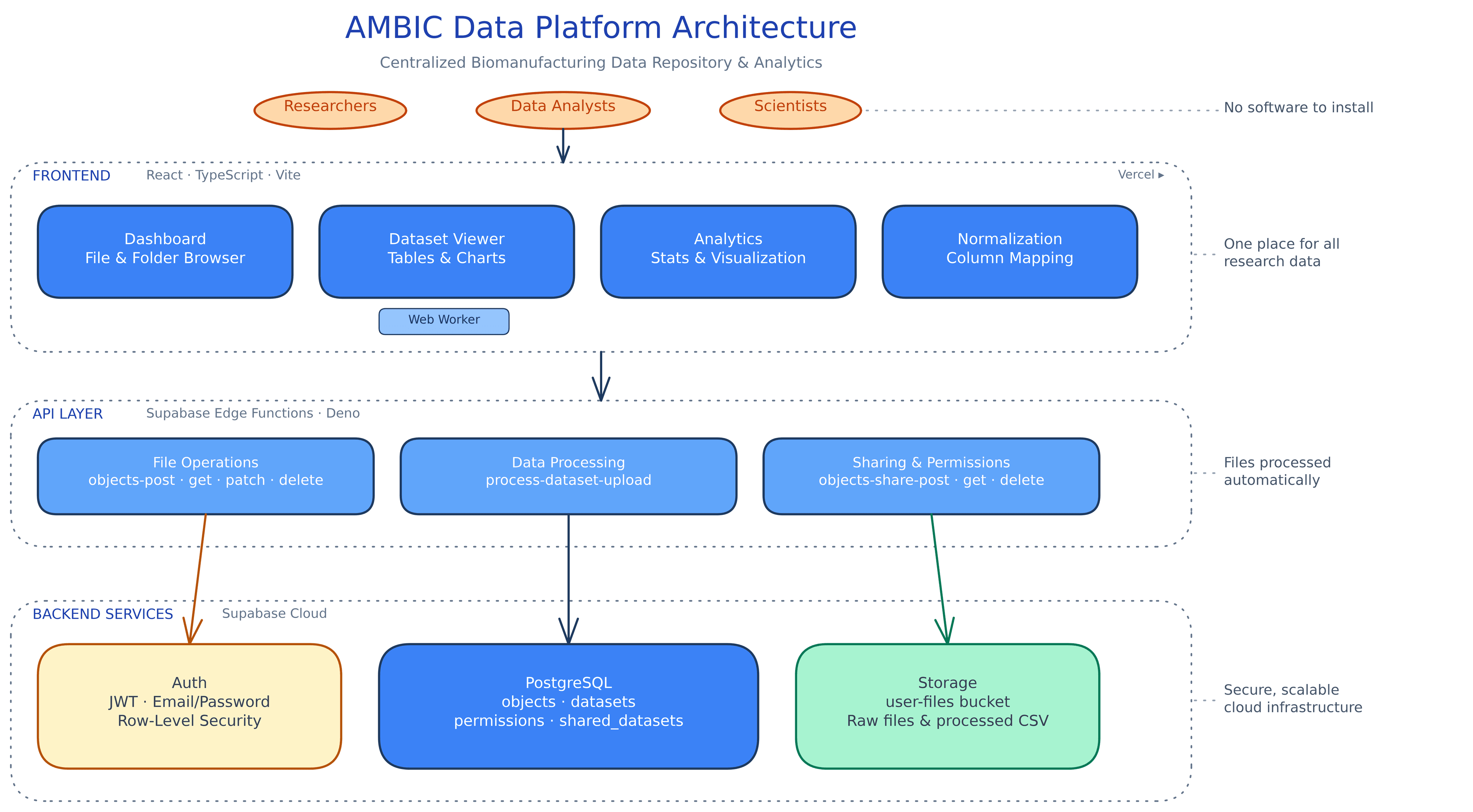
Architecture
Frontend is React + TypeScript with shadcn/ui and Recharts for interactive visualization. Backend runs on Supabase: PostgreSQL for structured data, Deno Edge Functions for file operations, object storage for raw uploads.
Stack
| Layer | Detail |
|---|---|
| Frontend | React 18, TypeScript, Vite (SWC), Tailwind CSS, shadcn/ui, Recharts |
| Tables | TanStack Table + TanStack Virtual, 2D row + column virtualization for 3,000+ column datasets |
| Backend | Supabase: PostgreSQL + Deno Edge Functions + object storage |
| Data parsing | Papaparse (CSV), ExcelJS (XLSX), native JSON |
| State | React Query (server), React Context (auth), URL params (navigation) |
| Auth | Supabase Auth (email/password) + MVP bypass token (internal testing) |
| Security | Row-level security on all tables; %/content/% path pattern for storage RLS |
| Edge Functions | Deno-based file CRUD, dataset processing, sharing operations |
Key Technical Challenges
- Wide datasets. Some biomanufacturing datasets have 3,000+ columns. The table component uses 2D virtualization (both row and column) so only the visible viewport renders. Without this, initial load would freeze the browser.
- Schema normalization with provenance. Researchers map source columns to a standardized target schema. Every normalized dataset maintains a link back to its source, preserving the full transformation history.
- Hierarchical file management. Folder nesting, dataset linking, and three-tier access control (owner, editor, viewer). The Supabase RLS policies for nested storage paths required careful attention, particularly around the
%/content/%path pattern matching.
Data Visualization
Interactive time series charts, data exploration dashboards, annotations, and dataset sharing with access control. The upload and normalization flow was designed to match how researchers already think about their experiments, not how the underlying database models the data.