DAEMON
Multi-agent LLM system coupled with mechanistic ODE simulations for autonomous scientific discovery. Agents explore parameter spaces on digital twins and produce human-readable hypotheses with Bayesian evidence tracking.
Context
Lyophilization (freeze-drying) of LNP/mRNA formulations is a process with a large parameter space, expensive lab runs, and poorly understood process-quality relationships. Pure ML approaches can fit the data but produce opaque models. The goal with DAEMON was to build a system that generates explicit, testable hypotheses rather than black-box predictions.
DAEMON stands for Discovery Agents Exploring Mechanistic ODE Networks. It was developed at the University of Massachusetts Lowell Department of Chemical Engineering.
Architecture
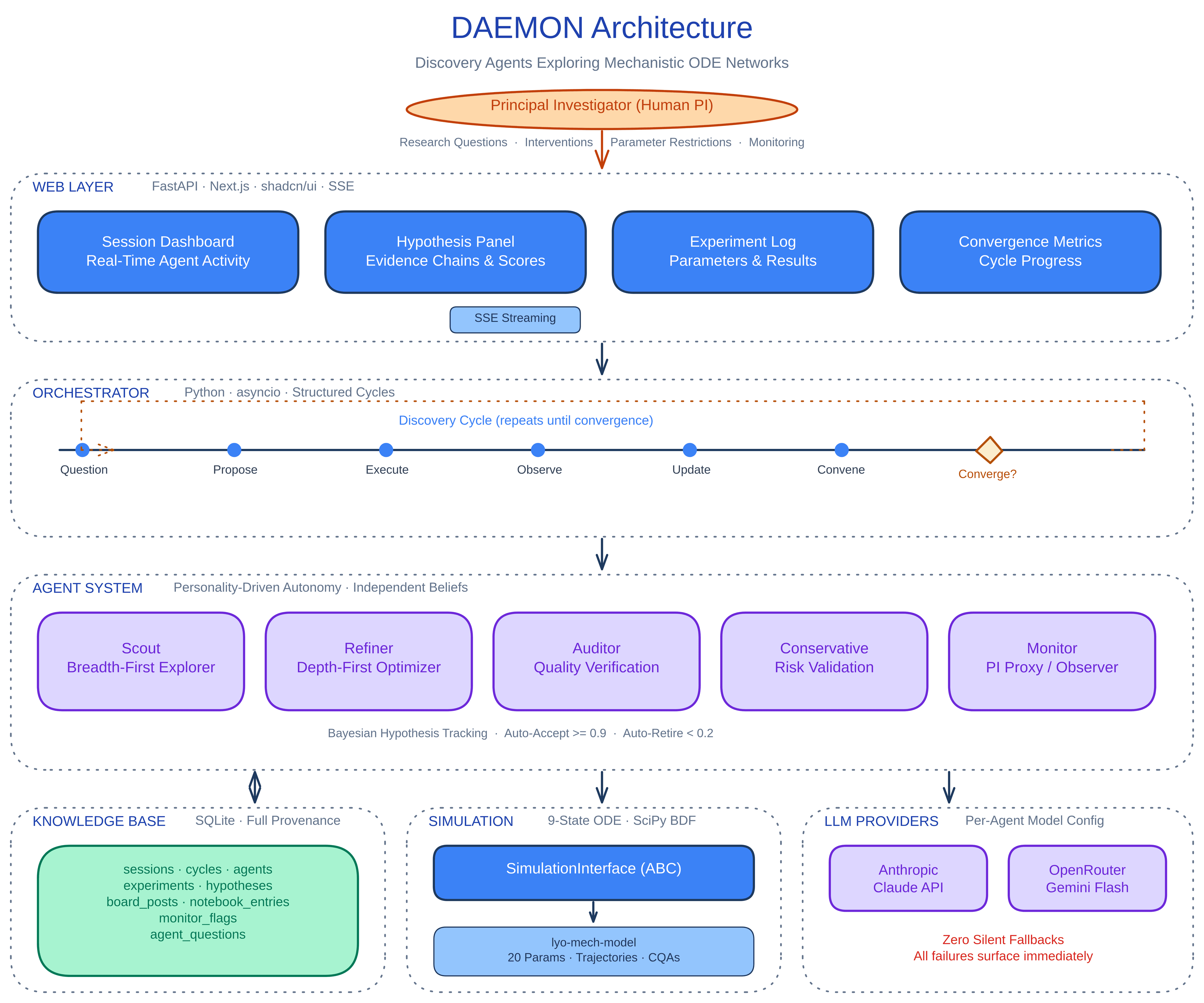
Five specialized agents (Scout, Refiner, Auditor, Conservative, Monitor) each maintain independent belief states and use different scientific reasoning strategies to propose experiments. They share a SQLite-backed knowledge base where hypotheses are tracked with Bayesian confidence scoring. The agents cooperate rather than compete: they review each other's conclusions, identify knowledge gaps, and flag contradictory evidence before the system declares convergence.
The simulation backend is a 9-state ODE model of the lyophilization process, tracking product temperature, ice thickness, chamber pressure, moisture, potency, cake resistance, encapsulation efficiency, RNA integrity, and particle size. Agents design and run experiments on this digital twin, then interpret the results against their current hypothesis set.
Monitoring Interface
The web layer is FastAPI (backend) + Next.js with shadcn/ui (frontend). Sessions stream in real time over SSE so a human PI can observe the discovery process and intervene when needed. The dashboard shows live agent activity, hypothesis evolution, experiment logs, and convergence metrics.
Stack
| Layer | Detail |
|---|---|
| Agents / Orchestrator | Python, Anthropic API, OpenRouter |
| Simulation | 9-state ODE kernel (lyo-mech-model submodule), SciPy solve_ivp, BDF method |
| Knowledge Base | SQLite, Bayesian hypothesis tracking, auto-accept >= 0.9, auto-retire < 0.2 |
| Backend | FastAPI, SSE streaming, asyncio with thread-offloaded SQLite |
| Frontend | Next.js 14, shadcn/ui, Tailwind CSS, Recharts |
| Agent Roles | Scout (exploration), Refiner (optimization), Auditor (verification), Conservative (risk), Monitor (convergence) |
| Reliability | Zero silent fallbacks. All LLM and simulation failures surface immediately. |
Design Decisions
This system is intended for pharmaceutical manufacturing contexts, so reliability constraints are strict. There are no silent fallbacks anywhere in the stack. If an LLM call fails or the simulation returns an error, the system surfaces it immediately rather than degrading to a heuristic. In life-critical applications, silent degradation is worse than a visible failure.
The discovery loop follows a structured cycle: propose, execute, observe, update, converge. Each cycle produces traceable artifacts. A researcher can read any hypothesis, follow its evidence chain back to specific simulation runs, and evaluate whether the reasoning holds.